Researchers can now investigate aneuploidy, a hallmark of cancer, using CRISPR
Research Paper
Nazario Bosco et al., KaryoCreate: A CRISPR-based technology to study chromosome-specific aneuploidy by targeting human centromeres. Cell. 186, 1985-2001.e19 (2023).Background and Scientific Question
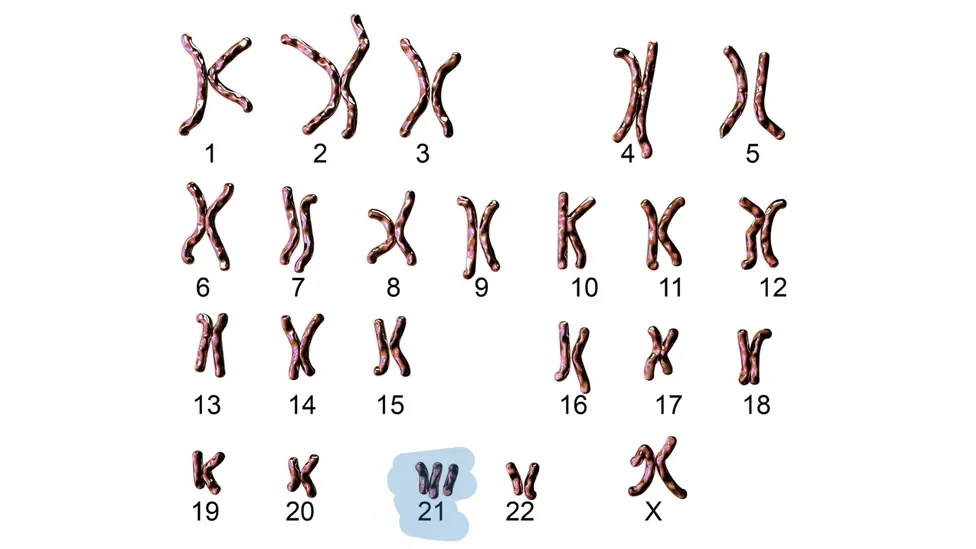
Humans have 23 pairs of chromosomes in their cells. The DNA sequence on these chromosomes contains genes encoding for proteins that keep the cell healthy. The gain or loss of chromosomes in cells is called aneuploidy. Aneuploidy is associated with many diseases including Down’s syndrome (gain of an additional copy of chromosome 21) and cancer 12. Aneuploidy in specific chromosomes is associated with specific cancers. For example, chromosome 3 is associated with some types of lung cancers 3.
To clearly study this association between aneuploidy of specific chromosomes and diseases, specific experimental models are required. However, these models are difficult to create. In today’s research paper, the authors address this problem by developing a new CRISPR-based technology to create specific aneuploidy models easily.
Experiments and Results
The researchers developed a CRISPR gene-editing-based approach to generate aneuploidy of specific chromosomes in cells. Let me first briefly describe the CRISPR gene-editing process.
In the CRISPR gene-editing process, there are 4 main components:
- The genome that you want to edit.
- A Cas9 protein that can cut the genome.
- A guide RNA that can direct the Cas9 protein to the correct region on the genome.
- A template DNA with an edited DNA sequence that can be copied into the cut genome.
Variations of these components can be used to get different results. For example, one can modify the Cas9 such that it does not cut. Such a Cas9 protein is called dCas9. This dCas9 can now be used as a vehicle to carry other proteins to a site on our genome.
In today’s research paper, the authors used a dCas9-based CRISPR approach. They tagged the dCas9 protein with a mutant form of the kinetochore protein KNL1. This mutant KNL1 can piggyback on the dCas9 and reach the chromosomes targeted by chromosome-specific guide RNAs. When mutant KLN1 reaches the chromosome, it causes the missegregation of that chromosome. This leads to aneuploidy. Therefore, a model of aneuploidy for a specific chromosome has been created.
The authors called this technology KaryoCreate (karyotype CRISPR-engineered aneuploidy technology).
Using this approach, the authors experimentally validated aneuploidy in 15 chromosomes in cultured human cells in the lab. They were able to observe chromosome segregation defects in these cells.
To further test their technology, they used their approach in colon cancer cells. They demonstrated that the loss of an arm of the q arm of chromosome 18 in these cancer cells promoted resistance to inhibitors of cell growth signals. Resistance to growth stop signals is a hallmark of cancer.
Conclusions and Outlook
Researchers developed a new CRISPR-based technology to generate aneuploidy models in human cells. These cells can be used to study aneuploidy and its association with cancer and other diseases. The researchers demonstrate an association with colon cancer. They showed that aneuploidy in colon cancer cells is associated with developing resistance to growth stop signals.
In the future, more research studies may use KaryoCreate to further understand aneuploidy. It may also uncover more associations between aneuploidy and human diseases.
References
- Stylianos E. Antonarakis et al., Down syndrome. Nature Reviews Disease Primers. 6, (2020).
- Uri Ben-David and Angelika Amon, Context is everything: aneuploidy in cancer. Nature Reviews Genetics. 21, 44-62 (2019).
- Alison M. Taylor et al., Genomic and Functional Approaches to Understanding Cancer Aneuploidy. Cancer Cell. 33, 676-689.e3 (2018).
